Dfttest
2D/3D frequency domain denoiser using Discrete Fourier transform (DFT)
| Abstract | |
|---|---|
| Author | tritical, cretindesalpes, DJATOM, pinterf |
| Version | v1.9.7 |
| Download | dfttest-v1.9.7.7z |
| Category | Spatio-Temporal Denoisers |
| License | GPLv2 |
| Discussion | Doom9 Thread, Update |
Requirements
- Supported color formats: Y8, YV12, YV16, YV24, YV411
- AviSynth+: all planar formats (8/10/12/14/16/32-bit, Y/YUV/RGB) are supported.
Runtime dependencies
The following are required, dfttest will not run or load without them.
- FFTW 3.3.5 (
fftw-3.3.5-dll32.ziporfftw-3.3.5-dll64.zip)
- *** 32-bit libfftw3f-3.dll needs to be in the search path (C:\Windows\SysWOW64 64-bit OS or C:\windows\system32 32-bit OS)
- *** 64-bit libfftw3f-3.dll needs to be in the search path (C:\windows\system32 64-bit OS)
Quick start
Denoising an 8-bit source
- Default options - moderate denoising (sigma) with moderate temporal filtering (tbsize):
dfttest(sigma=16, tbsize=5)
- Light denoising with less temporal filtering:
dfttest(sigma=6, tbsize=3)
- Light denoising with no temporal filtering:
dfttest(sigma=6, tbsize=1)
- sigma can be anywhere from 1.0 to 256.0 and beyond; denoising "strength" seems proportional to the square root of sigma.
- tbsize (temporal filter range) must be an odd number: 1, 3, 5, 7 ...etc
Denoising a high bit depth source
dfttest can accept a high bit depth (Stack16) source and return either 16bit (lsb=true) or 8bit (lsb=false).
- Strong denoising with no temporal filtering; convert output to 8bit
dfttest(sigma=64, tbsize=1, lsb_in=true, lsb=false)
- Same as above, but adding dither:
dfttest(sigma=64, tbsize=1, lsb_in=true, lsb=false, dither=1)
- dither=1 should combat any banding introduced by dfttest's quantization, but probably won't help banding in the source.
- dither=2 or higher adds random noise to combat banding in the source.
Syntax and Parameters
dfttest(clip clip [, bool Y, bool U, bool V, int ftype, float sigma, float sigma2,
float pmin, float pmax, int sbsize, int smode, int sosize, int tbsize,
int tmode, int tosize, int swin, int twin, float sbeta, float tbeta,
bool zmean, string sfile, string sfile2, string pminfile, string pmaxfile,
float f0beta, string nfile, int threads, int opt, string nstring, string sstring,
string ssx, string ssy, string sst, int dither, bool lsb, bool lsb_in, bool quiet ] )
clip clip =
- Source clip.
bool Y, U, V = true
- If true, the corresponding plane is processed. Otherwise, it is copied through to the output image as it is.
int ftype = 0
- Controls the filter type. Possible settings are:
ftype Filter Type 0 generalized wiener filter - mult = max((psd-sigma)/psd, 0)^f0beta
1 hard threshold - mult = psd < sigma ? 0.0 : 1.0
2 multiplier - mult = sigma
3 multiplier switched based on psd* value - mult = (psd >= pmin && psd <= pmax) ? sigma : sigma2
4 multiplier modified based on psd* value and range - mult = sigma * v((psd*pmax)/((psd+pmin)*(psd+pmax)))
- The real and imaginary parts of each complex DFT coefficient are multiplied by the corresponding mult value.
- * psd here means the power spectrum distribution (signal magnitude squared; real*real + imag*imag)[citation needed]
- The real and imaginary parts of each complex DFT coefficient are multiplied by the corresponding mult value.
float sigma, sigma2 = 16.0
- Value of sigma and sigma2 (used as described in ftype description).
- If using sfile or sstring then sigma is ignored.
- If using sfile2 then sigma2 is ignored.
- Value of sigma and sigma2 (used as described in ftype description).
NOTE: Starting in v1.5, these values are normalized based on the non-coherent power gain of the window when ftype<2. That is to say that for ftype<2, where sigma/sigma2 correspond to power, they are now independent of the window size and windowing function used, and that they directly correspond to power. For convenience, the normalization factor is output using OutputDebugString() when the filter loads. To convert between old and new sigma values, simply multiply the pre-v1.5 sigma value by the scaling factor. This scaling is also applied to values loaded from sfile/sfile2 files.
float pmin, pmax = 0.0, 500.0
- Used as described in the ftype description.
- If using pminfile then pmin is ignored.
- If using pmaxfile then pmax is ignored.
- Used as described in the ftype description.
NOTE: Starting in v1.5, these values are normalized based on the non-coherent power gain of the window. They are now independent of the window size and windowing function used, and directly correspond to power. For convenience, the normalization factor is output using OutputDebugString() when the filter loads. To convert between old and new pmin/pmax values, simply multiply the pre-v1.5 values by the scaling factor. This scaling is also applied to values loaded from pmin/pmax files.
int sbsize = 12
- Sets the length of the sides of the spatial window. Must be 1 or greater. Must be odd if using smode=0.
int smode = 1
- Sets the mode for spatial operation. There are two possible settings:
smode Operation 0 Process every pixel independently: center the spatial window on the current pixel, filter, move to the next pixel, repeat. Spatial overlapping sosize not used. 1 Process the spatial dimension in blocks of sbsize. Spatial overlapping is set from sosize.
int sosize = 9
- Sets the spatial overlap amount. Must be in the range 0 to sbsize-1 (inclusive).
- If sosize is greater than sbsize/2, then sbsize % (sbsize-sosize) must equal 0.
- In other words, overlap greater than 50% requires that sbsize-sosize be a divisor of sbsize.
- Sets the spatial overlap amount. Must be in the range 0 to sbsize-1 (inclusive).
int tbsize = 5
- Sets the length of the temporal dimension (i.e. number of frames). Must be at least 1. Must be odd if using tmode=0.
int tmode = 0
- Sets the mode for temporal operation. There are two possible settings:
tmode Operation 0 Process every frame independently: center the temporal window on the current frame, filter, move to the next frame, repeat. Temporal overlapping tosize not used. 1 Process the temporal dimension in blocks of tbsize. Temporal overlapping set from tosize.
int tosize = 0
- Sets the temporal overlap amount. Must be in the range 0 to tbsize-1 (inclusive).
- If tosize is greater than (tbsize/2), then tbsize%(tbsize-tosize) must equal 0.
- In other words, overlap greater than 50% requires that tbsize-tosize be a divisor of tbsize.
- Sets the temporal overlap amount. Must be in the range 0 to tbsize-1 (inclusive).
int swin, twin = 0, 7
- Sets the type of analysis/synthesis window to be used for spatial (swin) and temporal (twin) processing. Possible settings:
swin/twin Window 0 hanning 1 hamming 2 blackman 3 4-term blackman-harris 4 kaiser-bessel 5 7-term blackman-harris 6 flat top 7 rectangular 8 Bartlett 9 Bartlett-Hann 10 Nuttall 11 Blackman-Nuttall
float sbeta, tbeta = 2.5
- Sets the beta value for kaiser-bessel window type.
- sbeta goes with swin, tbeta goes with twin.
- Not used unless the corresponding window value is set to 4.
- Sets the beta value for kaiser-bessel window type.
bool zmean = true
- Controls whether the window mean is subtracted out (zeroed) prior to filtering in the frequency domain.
string sfile = ""
- Specifies an input file listing sigma values for each DFT coefficient.
- There can be multiple lines with multiple coefficients per line.
- Separate coefficients on the same line using ',' or ' '.
- Placing a '#' at the beginning of a line will cause that line to be ignored.
- The coefficients are read from the file in left to right, top to bottom order.
- Specifies an input file listing sigma values for each DFT coefficient.
- The DFT transform results in tbsize*sbsize*(sbsize/2+1) coefficients. You must give these many sigma values in the sfile. Assuming 2D (tbsize=1), and sbsize=8 (i.e. 8x8 window). The resulting transform has 40 coefficients, organized as follows:
0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39
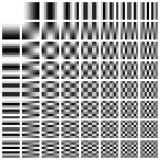
- The following graphic from Wikipedia:DCT shows the frequency arrangement:
- The numbers here specify which sigma value from the sfile corresponds to that DFT coefficient.
- The DC (frequency=0) coefficient is in the upper left. The top row corresponds to purely horizontal frequencies, and the frequencies increase from left to right. In this example '4' corresponds to the highest horizontal frequency.
- The left-most column corresponds to purely vertical frequencies, but the highest frequency is at the sbsize/2 row (assuming the numbering starts at 0)... in this case '20' corresponds to the highest vertical frequency. The frequencies then decrease from sbsize/2 to sbsize-1. Basically, the first sbsize/2 rows correspond to the positive frequencies and the last sbsize/2-1 rows correspond to the negative frequencies.
- In the 8x8 case, the single highest frequency is located at '24'.
- In the case that tbsize>1, the first set of (sbsize/2+1)*sbsize coefficients correspond to the lowest frequencies temporally (with the relations described for the 2D case holding within that set) and the frequencies increase temporally from set to set up to the tbsize/2 set. The frequencies then decrease from there to the tbsize-1 set (again the positive vs negative frequencies as mentioned previously). If tbsize=3, you get 120 coefficients:
0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119
- The DC coefficient is still at '0'. The highest purely temporal frequency is at '40'. The highest overall frequency is at '64'.
string sfile2, pminfile, pmaxfile = ""
- Can be used to give different values of sigma2, pmin, and pmax for each DFT coefficient respectively.
- Entry and format is exactly the same as described in the sfile parameter description.
- If sfile2 is not given then the value of sigma2 is used for every coefficient.
- If pminfile is not given then the value of pmin is used for every coefficient.
- If pmaxfile is not given then the value of pmax is used for every coefficient.
- Can be used to give different values of sigma2, pmin, and pmax for each DFT coefficient respectively.
float f0beta = 1.0
- Power term in ftype=0. The ftype=0 formula is:
- max((psd-sigma)/psd, 0)^f0beta
- For f0beta=1, this equation corresponds to the wiener filter with spectral subtraction as the estimate of the signal power.
- For f0beta=0.5, the equation corresponds to spectral subtraction.
- The 1.0 and 0.5 cases are separated from the general routine in the code to allow for fast operation.
- Other values will result in the general routine being used, which has to perform a pow() computation, and is therefore much slower.
- For f0beta=1, this equation corresponds to the wiener filter with spectral subtraction as the estimate of the signal power.
string nfile = ""
- When ftype<2, an nfile can be used to specify block locations in the video from which dfttest will estimate the noise power spectrum (sigma) to be used for filtering.
- When the noise to be removed is not white (i.e. doesn't have a flat power spectrum), specifying only a single sigma value is not adequate. Prior to v1.5, using dfttest in such cases meant you would have to figure out the noise spectrum on your own, and then use an sfile to input the sigma values. Now dfttest can perform the task of estimating the noise spectrum.
- The nfile should list locations in the video that consist of noise on a flat background, one entry per line. The line syntax is:
- frame_number,plane,ypos,xpos
- set plane to 0 for the Y plane, 1 for the U plane, or 2 for the V plane.
- set ypos & xpos to the upper left position of the block [in pixels]
- (0,0 is the upper left of the frame)
- for example,
- 0,0,20,20
- dfttest positions a window (of the type defined by sbsize/tbsize/swin/twin) at the specified location, and estimates the power using fft magnitude^2. When tbsize>1, frame_number specifies the first frame of the temporal block. Make sure that the window size is large enough to capture the full noise pattern.
- If you list multiple blocks (multiple lines in the nfile), then the estimates obtained at each block are averaged to form the final estimate. Having more block locations to use lowers the variance of the estimate. The more block locations you specify the closer the true noise spectrum will be estimated, resulting in better denoising. When listing multiple block locations, it is best/preferred if the locations do not overlap.
- Typically, subtracting out the noise power spectrum is not adequate because it is only the average. In any one block the noise spectrum has the potential to exceed the average in a frequency bin. Therefore, one typically over-subtracts based on some multiple of the noise spectrum (usually in the range of 3-8). The default over-subtraction factor is 5 if ftype=0 and 7 if ftype=1. If you want to use another value, then on some line in the nfile put the following:
- a=over_subtraction_factor
- for example,
- a=3.5
- To comment out a line in an nfile (have it be ignored), place a '#' at the beginning of the line.
- An example:
avisource("noisy_source.avi")
dfttest(f0beta=0.5, U=false, V=false, nfile="nfile.txt")
- Here, dfttest is filtering the Y plane only, using default settings except for f0beta=0.5, resulting in spectral subtraction instead of Wiener filtering. nfile is listing locations of only noise, and has the following lines:
0,0,20,40 5,0,100,380 14,0,400,100 a=5.2
- The first line specifies a block from frame 0, plane 0 (Y), at x,y location (40,20). The next two lines specify two additional blocks. The estimate from all three blocks will be averaged. On the last line, the over-subtraction factor is set to 5.2.
- When using an nfile, the estimated noise spectrum is output to "noise_spectrum-date_string.txt", located in the current directory. It lists the power of each DFT coefficient. The layout is the same as explained in the sfile description. The average noise power is also calculated. As of v1.7, this file is compatible (can be used) with sfile.
int threads = 0
- Sets the number of threads used for processing. If set to 0, then threads is set equal to the number of detected processors.
int opt = 0
- Sets which CPU optimizations are used. Possibly use for debug purposes, e.g. try C version intentionally. Possible settings:
opt CPU Optimizations 0 auto detect 1 C routines 2 SSE/SSE2 routines 3 AVX routines 4 AVX2 routines
string nstring = ""
- Same functionality as nfile, but allows entering window locations directly in the script instead of creating a separate file. The list of frame/plane/ypos/xpos quadruples is stored as a string with each quadruple separated by a space.
- Example - If you use an nfile that looks like:
a=4.0 35,0,45,68 28,0,23,87
- You can use the following nstring and get the same result:
nstring="a:4.0 35,0,45,68 28,0,23,87"
- The one restriction is that the over-subtraction factor (a:x.x) must be the first entry in the string, as opposed to nfiles where the a=x.x can be placed anywhere. If it is not supplied, then the same default over-subtraction factor is used as is used for the nfile option.
string sstring, ssx, ssy, sst = ""
- Used to specify functions of sigma based on frequency.
- If you want sigma to vary based on frequency, then use 'sstring' instead of sigma. sstring allows you to enter values of sigma for different normalized [0.0,1.0] frequency locations.
- Values for locations between the ones you explicitly specify are computed via linear interpolation. The frequency range, which is dependent on sbsize/tbsize, is normalized to [0.0,1.0] with 0.0 being the lowest frequency and 1.0 being the highest frequency.
- You MUST specify sigma values for those end point locations (0.0 and 1.0). You can specify as many other locations as you wish, and they don't have to be in any particular order.
- Each frequency/sigma pair is given as f.f:s.s. The list of frequency/sigma pairs is saved as a string, with each pair separated by a space.
- Used to specify functions of sigma based on frequency.
- For example, if you want a linear ramp of sigma from 1.0 for the lowest frequency to 10.0 for the highest frequency use:
- sstring = "0.0:1.0 1.0:10.0"
- "0.0:1.0" → sigma= 1.0 at frequency 0.0
- "1.0:10.0" → sigma=10.0 at frequency 1.0
- sigma values for frequencies between 0.0 and 1.0 will be computed via linear interpolation.
- Or if you want a band-stop filter that passes low and high frequencies (filters middle frequencies) use something like:
- sstring = "0.0:0.0 0.15:10.0 0.85:10.0 1.0:0.0"
- To help visualize the process, the resulting filter spectrum is output to "filter_spectrum-date_string.txt" using the same format as the "noise_spectrum-date_string.txt" file that is output by the nfile/nstring options. The format of this file is compatible with sfile input.
- There are two methods for computing sigma values for a given frequency bin based on sstring. The first computes the normalized frequency location of each dimension (horizontal, vertical & temporal), interpolates sigma for each of those dimensions, and then multiples the individual sigmas to obtain the final sigma value. So that everything scales correctly, all sigma values entered in sstring are first raised to the 1/#_dimensions power before perform performing linear interpolation and multiplying. The second method (based on FFT3DFilter's system) works by computing a single location from the separate dimension locations (x,y,z) as:
- new = sqrt((x*x+y*y+z*z)/3.0)
- sigma is then interpolated to this location. By default the first system is used. To use the second system simply put a '$' sign at the beginning of sstring as shown below:
- sstring = "$ 0.0:1.0 1.0:10.0"
ssx / ssy / sst explanation
'sstring' breaks the 1D (sbsize=1), 2D (for tbsize=1), or 3D (for sbsize>1 and tbsize>1) frequency spectrum into chunks by normalizing each dimension to [0.0,1.0]... i.e. the frequency range [0.0,0.25] is a cube covering the first 1/4 of each dimension. This works fine if you want to treat all dimensions the same in terms of how sigma should vary. However, if you wanted to ramp sigma based only on temporal frequency or horizontal frequency, this is too limited. This is where 'ssx'/'ssy'/'sst' come in!
ssx/ssy/sst allow you to specify sigma as a function of horizontal ('ssx'), vertical ('ssy'), and temporal ('sst') frequency only. The syntax is exactly the same as that of sstring. To get the final sigma value for a frequency location, the three separate values (one for each dimension) are computed and then multiplied together. As with sstring the sigma values are first raised to the 1/#_dimensions power before performing linear interpolation and multiplying. If you don't specify all three strings, then a flat function equal to sigma is used for the missing dimensions. For dimensions of size one (the spatial dimensions if sbsize=1 or the temporal dimension for tbsize=1) the corresponding string is ignored.
For example:
ssx="0.0:1.0 1.0:10.0",ssy="0.0:1.0 1.0:10.0",sst="0.0:1.0 1.0:10.0"
will give the same result as
sstring="0.0:1.0 1.0:10.0"
Or if you want to ramp sigma based on temporal frequency:
sigma=10.0,sst="0.0:1.0 1.0:10.0"
This will use 10.0 for the horizontal/vertical dimensions, and ramp sigma from 1.0 to 10.0 in the temporal dimension.
- If sstring is specified, it takes precedence over ssx/ssy/sst. Again, the "filter_spectrum-date_string.txt" output file is helpful in visualizing the result.
int dither = 0
- Controls whether dithering is performed when converting from float to unsigned char for output. Internally dfttest works on floating point values. For output the result must be quantized back to unsigned char values. Prior to v1.8 this was always done by simply rounding. Possible settings:
dither Operation 0 No dithering (same as v1.7 and prior) 1 Floyd-Steinberg dithering 2-100 Floyd-Steinberg dithering with increasing amounts of uniform random noise added prior to the dithering process
- Obviously dither=0 is the fastest, and dither=1 is slightly faster than dither>=2 due to not having to generate a random number for every pixel. However, this part doesn't take much time compared to the actual filtering operation. dither=1 should combat any banding introduced by dfttest's quantization, but probably wont help banding in the source. dither>=2 can combat banding in the source.
bool lsb = false
- Note: since v1.9.6 all Avisynth+ high bit depth formats are supported, so this option is deprecated.
- When lsb=true, dfttest outputs 16-bit pixel components by separating the most significant bytes (MSB) and the least significant bytes (LSB). The top part of the frame contains the MSB of all pixels and the bottom part their LSB. Therefore the output frame height is doubled. Use this if you want to perform the dithering later, with a separate tool.
bool lsb_in = false
- Note: since v1.9.6 all Avisynth+ high bit depth formats are supported, so this option is deprecated.
- When lsb=true, the input is supposed to have 16-bit pixel components, of the same format as the output given with lsb=true. The sigma scale remains relative to the MSB, meaning that a given value will have the same visual results with 16-bit and 8-bit clips.
bool quiet = true
- If quiet=false, creates a filter spectrum file when sigma is specified with sstring or ssx
- If quiet=true, no file is created.
Examples
- TODO
Changelog
| Version | Date | Description |
|---|---|---|
| v1.9.7 | 2021-10-28 |
+ pass Avisynth+ frame properties |
| v1.9.6 | 2020-03-24 |
+ Full high bit depth support under Avisynth+ |
| v1.9.4.3 | 2018-10-14 |
+ x64 version |
| v1.9.4 | 2013-08-04 |
+ Compatible the new Avisynth 2.6 colorspaces, except Y8. |
| v1.9.3 | 2012-04-20 |
- Does no longer issue a tbsize-related error with null-length clips. |
| v1.9.2 | 2012-03-23 |
- The quiet parameter is not true by default. |
| v1.9.1 | 2012-03-11 |
- Fixed a stupid regression (from v1.8 mod16a) on the dither parameter. |
| v1.9 | 2011-11-28 |
+ Added the quiet parameter to deactivate the filter spectrum output. |
| v1.8 mod16b | 2011-05-12 |
+ Added the lsb_in parameter to input 16 bit data. |
| v1.8 mod16a | 2010-06-26 |
+ Added the lsb parameter to output 16 bit data. |
| v1.8 | 2010-06-22 |
+ added dither parameter and functionality |
| v1.7 | 2010-06-21 |
+ added nstring/sstring/ssx/ssy/sst parameters and functionality |
| v1.6 | 2009-06-04 |
- fixed window normalization causing tmode=0 to always result in a rectangular temporal window, and smode=0 to always result in a rectangular spatial window. |
| v1.5 | 2009-04-11 |
+ added f0beta in ftype=0 |
| v1.4 | 2009-04-06 |
- fix threading issue that could result in corrupted output |
| v1.3 | 2009-01-27 |
+ more assembly optimizations |
| v1.2 | 2009-01-24 |
+ added filter types 3/4 and corresponding parameters (sigma2,pmin,pmax, sfile2,pminfile,pmaxfile) |
| v1.1 | 2007-11-22 |
+ more sse optimizations |
| v1.0 | 2007-11-21 |
- initial release |
Archived Downloads
| Version | Download | Mirror |
|---|---|---|
| v1.9.4.3 | dfttest-1.9.4.3 | |
| v1.9.4 | dfttest-1.9.4.zip | dfttest-1.9.4.zip |
| v1.8.0 | dfttestv18.zip |
External Links
- GitHub - source code repository (v1.9.4.3 by DJATOM).
- GitHub - source code repository (v1.9.x by pinterf).
Back to External Filters ←